/Project Details
Simulation-Based Inference for the SWIFT Eye-Movement Model
Implemented a BayesFlow pipeline for a simplified SWIFT model to infer gaze-control parameters from simulated eye-tracking fixation sequences.
This project explores the SWIFT model of eye movements as a simulation-based inference problem. It focuses on how readers decide when to move their eyes and where to look next while processing written language.
The full SWIFT model is computationally intensive and has an intractable likelihood, so the project used a simplified SWIFT-style simulator inspired by Engbert and Rabe (2024). The goal was to make the model usable inside an amortized Bayesian inference workflow with BayesFlow.
I built a Python notebook that loads fixation sequences, filters very short fixations, creates sentence-level metadata, defines priors for gaze-control parameters, and simulates fixation duration, word position, saccade length, landing error, skip behavior, refixation behavior, skip rate, and refixation rate.
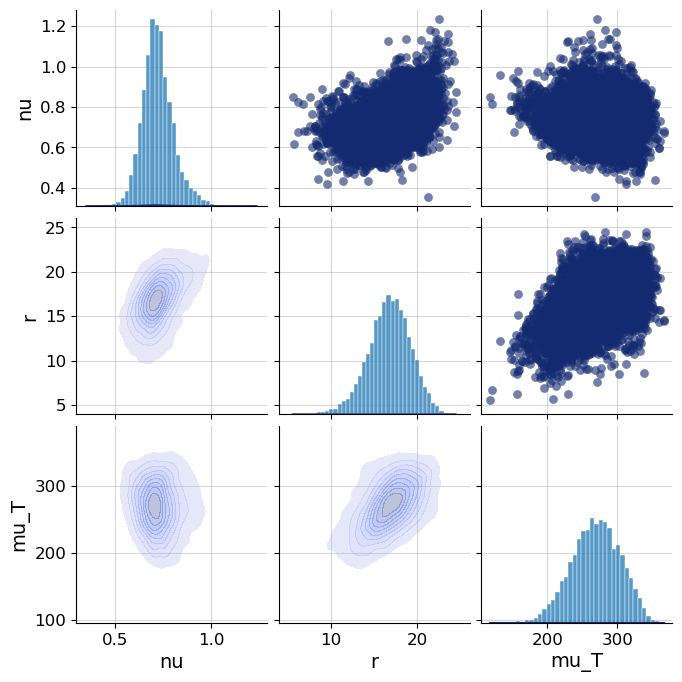
The inference target was a compact parameter vector including nu, r, mu_T, and eta. I connected the simulator to BayesFlow, built adapters to concatenate inference variables and summary variables, and trained neural inference workflows using FlowMatching, SetTransformer-style summaries, and later point-estimation experiments with TimeSeriesNetwork and PointInferenceNetwork.
Methodologically, the project is shaped by the modern Bayesian workflow around BayesFlow, amortized Bayesian inference, and the broader probabilistic modeling ecosystem associated with Prof. Dr. Paul Bürkner. It also builds on the simplified SWIFT tutorial by Engbert and Rabe (2024).
Highlights
- Implemented a simplified SWIFT-style simulator for fixation duration, saccade length, skip/refixation behavior, and landing error.
- Defined priors for nu, r, mu_T, and eta to represent gaze-control and reading-dynamics parameters.
- Connected custom simulation code to BayesFlow using simulator metadata, adapter transformations, and neural summary variables.
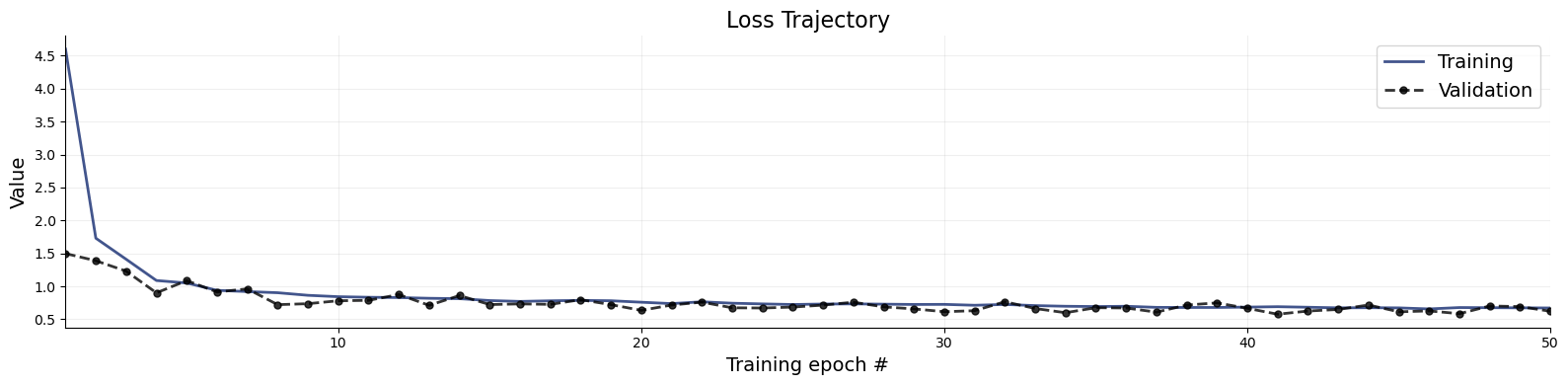
- Trained an online BayesFlow workflow for 50 epochs with 100 batches per epoch and monitored validation loss and default diagnostics.
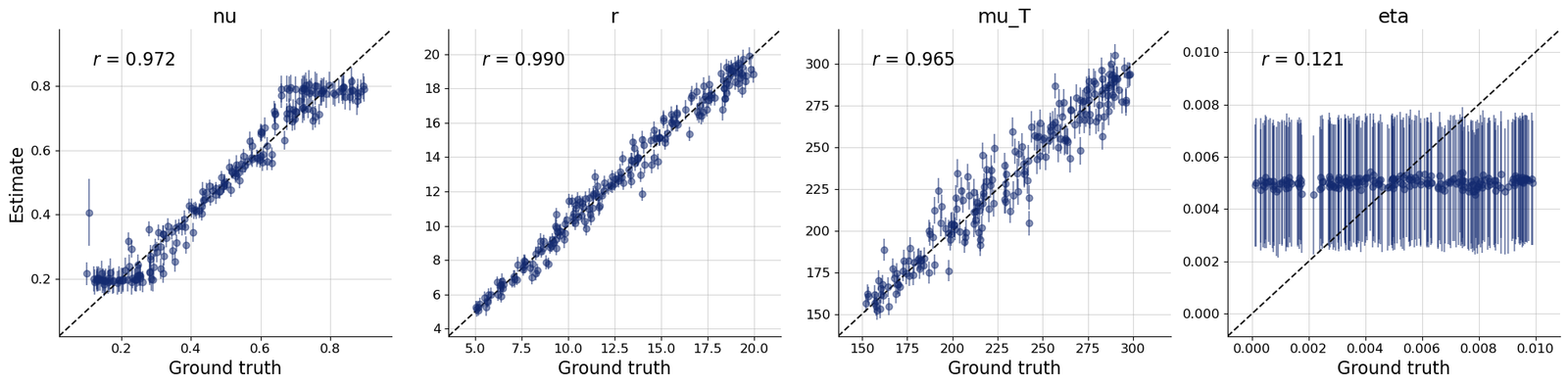
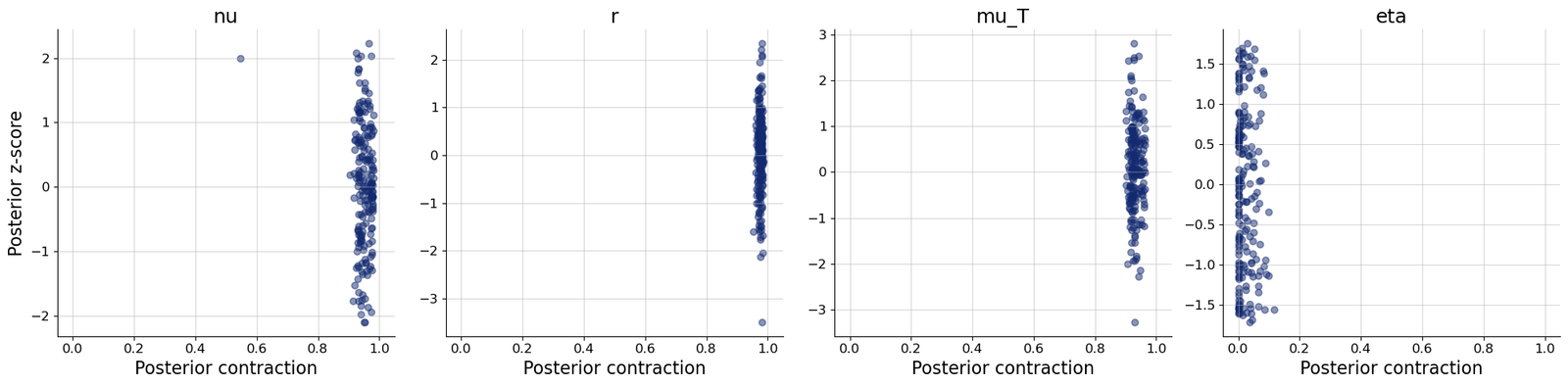
- Generated posterior samples and diagnostic plots including loss, recovery, calibration ECDF, and z-score contraction workflows.
- Explored fallback BayesFlow 2.x APIs with TimeSeriesNetwork, ContinuousApproximator, PointApproximator, and PointInferenceNetwork to handle package-version constraints.
Figures